We are proud to announce publication of a new research article in American Journal of Transplantation (AJT)!
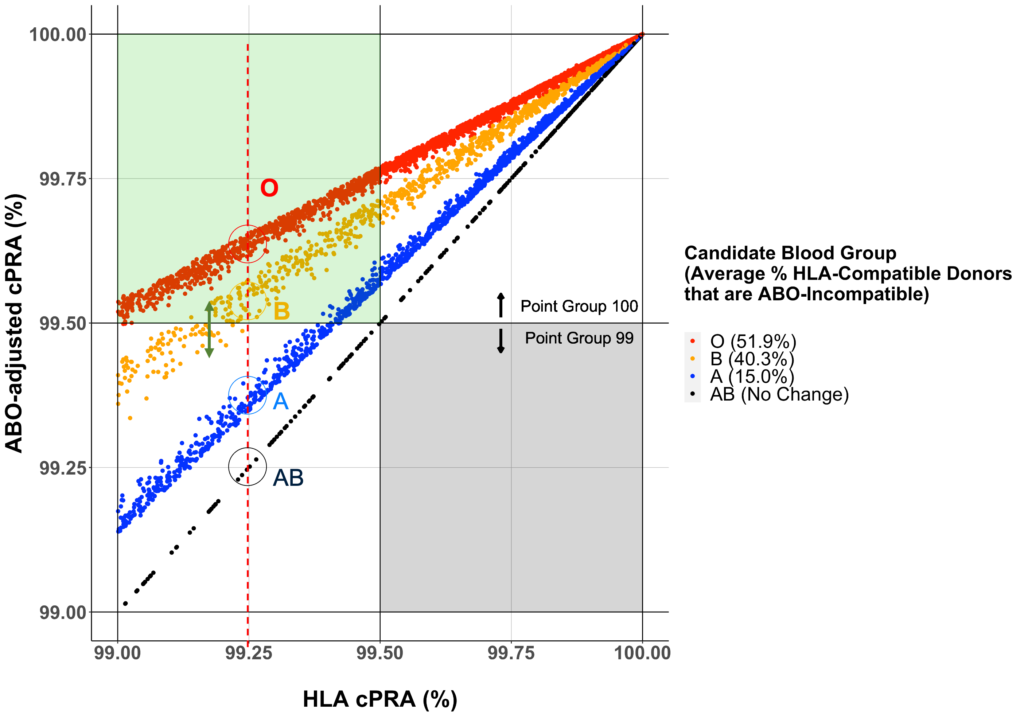
The paper validates a new approach to measuring antibody sensitization using a HLA typed reference panel of 6.59 million stem cell donors from National Marrow Donor Program (NMDP). The current United Network for Organ Sharing (UNOS) calculator uses 14,282 deceased donors and does not include two-field alleles or the DQA1 and DPB1 loci.
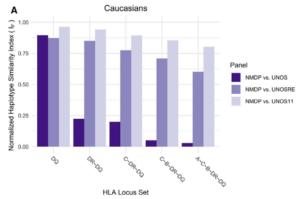 The figure shows a key result from validation of a NMDP reference panel for calculated panel reactive antibody (CPRA) calculations. We found to our surprise that in the Caucasian population that HLA haplotype frequencies differed dramatically between the NMDP and current UNOS panel. We were able to recover expected linkage disequilibrium values in the UNOS dataset by recomputing haplotype frequencies (UNOSRE) using Arlequin EM algorithm. By expanding the UNOS reference panel to 11 years of deceased donors, we achieved higher similarity, indicating that most of the remaining differences with the NMDP dataset are due to sampling error.
The figure shows a key result from validation of a NMDP reference panel for calculated panel reactive antibody (CPRA) calculations. We found to our surprise that in the Caucasian population that HLA haplotype frequencies differed dramatically between the NMDP and current UNOS panel. We were able to recover expected linkage disequilibrium values in the UNOS dataset by recomputing haplotype frequencies (UNOSRE) using Arlequin EM algorithm. By expanding the UNOS reference panel to 11 years of deceased donors, we achieved higher similarity, indicating that most of the remaining differences with the NMDP dataset are due to sampling error.
Based on this proof of concert, the UNOS Histocompatibility Committee is moving forward with this approach for updating the CPRA calculator based on NMDP data to include all classical HLA loci at high resolution. This will help ensure that highly sensitized patients are treated fairly and equitably in the kidney allocation system. Stay tuned for more news on this!
Kransdorf, E.P., Pando, M.J., Stewart, D., Lindblad, K., Bray, R., Murphey, C., Kaur, N., Patel, J.K., Kim, I., Zhang, X., Maiers, M., Kobashigawa, J.A., Gragert, L. Stem cell donor HLA typing improves CPRA in kidney allocation. American Journal of Transplantation (2020).
https://pubmed.ncbi.nlm.nih.gov/32558252/